|
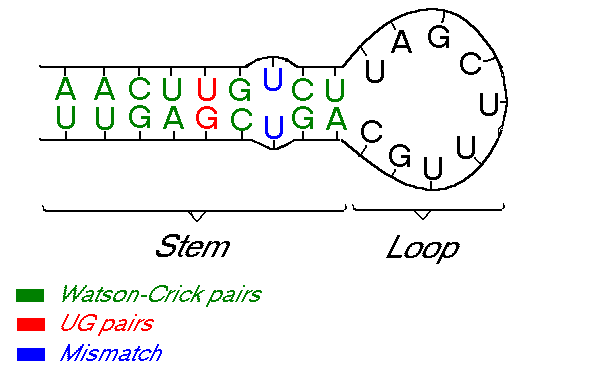
There are 16 possible base-pairings, however of these, only six (AU, GU, GC, UA, UG, CG) are stable enough to form actual base-pairs. The rest are called mismatches and occur at very low frequencies in helices. RNA molecules, such as ribosomal RNAs and transfer RNAs, have an important role. Their structure cannot easily be disrupted without impact on their function and lethal consequences and selection is acting to maintain the secondary structure. Yet, the primary structure of the stems (i.e., their nucleotide sequence) can still vary and in fact we observe that RNA helical regions are quite variable in sequence. The nature of the bases is not important and substitutions are possible as long as they preserve the secondary structure. One could model the evolution of stems using the DNA models described above but there may be a substantial bias in results because paired substitutions would seem far less probable than they are in reality (see Jow et al., 2002). Statistics become invalid and it can have an effect on inferred phylogenies.
The secondary structure is left unchanged when complementary substitutions occur in the DNA gene coding for the RNA molecule. The process can be a single step process (double substitution) or a two step process (two single substitutions). These two processes are descibed in the theory of compensatory substitutions section.